CRISPR-Cas9: An Evolutionary Technique for the Treatment of Breast and Cervical Carcinoma
Download
Abstract
CRISPR/Cas9 has transformed genome editing methods in several scientific domains, including the study of human cancer. Globally, breast cancer (BC) affects women more than any other kind of cancer and triple-negative breast cancer (TNBC) has the greatest fatality rate. The diagnosis and treatment of breast cancer have come a long way, yet many patients still experience metastases or relapses. One of the potential techniques for correcting faulty genes and curing different cancers is gene therapy. Certain mutations associated with cancer onset and development might be repaired in model organisms as a first step toward translational applications using the CRISPR/Cas9 technology. The CRISPR/Cas9 system, which consists of a single readily modified guide RNA (sgRNA) sequence coupled to a Cas9 nuclease, has transformed genome editing owing to its simplicity and effectiveness when compared to previous methods. CRISPR/Cas9 allows for the knockdown of over-expressed genes, the reversal of mutations in defected genes, and the remodeling of the regulatory environment to inhibit BC proliferation. CRISPR/Cas9-based treatment mixed with traditional chemotherapy has proven to be beneficial in combating drug resistance and tumor inhibition difficulties. The CRISPR cas9 system may edit many genes at the same time by generating numerous sgRNAs that target different genomic regions, considerably increasing its therapeutic potential and making it a unique technique for treating cervical lesions. This review focuses on the therapeutic applications of CRISPR/Cas9 in the treatment of cervical cancer and breast cancer.
Introduction
Numerous novel gene editing techniques have been discovered in the last 10 to 20 years and are currently being employed in a variety of applications. The CRISPR/ Cas9 system stood out among all of those techniques as a pioneering technique that allows the insertion, deletion, and correction of genetic material both in vivo and in vitro. The medical and health fields have been significantly transformed by this method [1]. Using a guided RNA to locate its target sequence, this method subsequently inserts or deletes a DNA segment with the aid of the Cas9 enzyme [2]. The natural repair systems of the cells, homology-directed repair (HDR) and non-homologous end joining (NHEJ), are triggered when the cleavage occurs [3, 4]. It is possible to insert donor DNA to serve as a template at the location of cleavage by using the right mechanism and design.
The second most frequent cancer in women to cause mortality worldwide is breast cancer [5, 6]. Breast cancer accounts for 28% of all new cancer cases [7]. It develops silently and is detected on routine screening. Both the initiation and development of breast cancer involve several signaling pathways [8]. Studies have revealed that the expression of the estrogen receptor (ER), progesterone receptor (PR), and human epidermal growth factor receptor 2 (HER2) is linked to a number of breast cancer subtypes [9]. Some molecular markers associated with breast cancer are represented in Figure 1.
Figure 1. Molecular Subtypes of Breast Cancer.
Adverse physical side effects, non-specificity and drug resistance are major problems even if there aren’t many medications licensed for the treatment of breast cancer [10, 11]. CRISPR/Cas9 technology is getting a lot of interest as research on breast cancer continues to be a hot topic.
CRISPR/Cas9 is the newest type of gene editing technology among all available techniques, yet research has shown that it is also the most effective one. This technique does have some drawbacks as well; however, researchers are striving to address these limitations [12]. The CRISPR-CAS system is valuable for studies on cancer, particularly breast cancer [13]. The therapeutic promise of the CRISPR/Cas9 system against breast and ovarian cancer is highlighted in this review. We will explain how the CRISPR/Cas9 system can be used to overcome drug resistance in breast and ovarian cancer. We’ll discuss about the CRISPR/Cas9 system’s limitations, advances, and the future prospects in the final section.
CRISPR-CAS: A gift from the nature
The eukaryotic genome contains millions of bases that are hard to modify, but efforts are being made to find new anticancer targets and design new therapeutic approaches. The natural DNA repair machinery is stimulated by double stranded bricks (DSBs) in the DNA, according to studies. By using non-homologous end joining (NHEJ), mutant sequence can be added to or removed from the cleavage site. However, homology directed repair (HDR) in animal models permits both knock-in and knock-out (Figure 2).
Figure 2. Genome Editing Using Natural Repair Mechanism of Host.
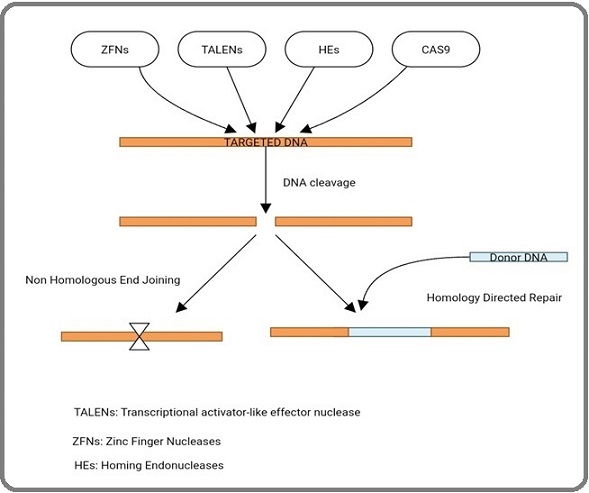
These investigations aided in the discovery of numerous novel gene editing tools [11].
Prokaryotes’ defense mechanism led to the discovery of the CRISPR/Cas9 system, which turned out to be a gift from nature. CRSIPR-CAS was first identified in the 1980s as a prokaryote defense system, but it wasn’t until 2012 that its potential for use as a gene editing system was investigated. Bacteria’s CRISPR-CAS system cleaves foreign genetic material and functions as an immune system. Using guided RNA and the CAS9 protein, the CRISPR/Cas9 system can induce double stranded bricks on the targeted sites of the genome [14, 15]. The CRISPR- Cas9 system needs a protospacer adjacent motif (PAM) next to the targeted region in order to recognize and cleave the DNA [16, 17]. Regarding the CRISPR-Cas9 mechanism, 60% of archaea and 40% of bacteria show minor differences [18]. The spacers in the CRISPR locus act as memory repositories and shield the host from attacks from the same invader over and over again.
CRISPR/Cas9 system is widely used in the breast cancer research due to its effectiveness. The effectiveness of the CRISPR/Cas9 system is due to its ease of use. In comparison to previous methods, the CRISPR/Cas9 system is simpler to construct and less expensive. The PAM requirement is a restriction, although this can be bypassed by employing different CAS proteins. The bases and methylation patterns can be altered with the help of enzymes like DNA methyltransferases (DNMT) and DNA deaminases. Such modifications can improve the therapeutic potential of the CRISPR/Cas9 system in the treatment of diseases like breast cancer. Some Targeted genes of different cell types in animals and vector used are being represented in Table 1.
| Sr. | Cell Type | Animal | Vector | Genes Targeted |
| 1 | Germline | Mouse | mRNA,sgRNA | Tet1, Tet2 |
| Rat | mRNA,sgRNA | Not Specific | ||
| Monkey | mRNA,sgRNA | Pparg, Rag1 | ||
| Mouse | Plasmid DNA | p53, pten, APC | ||
| 2 | Somatic | Mouse | AAV | Mecp2 |
| Mouse | sgRNA | Icam2 | ||
| Mouse | Lentivirus | APC, pten | ||
| Mouse | Adenovirus | Eml4-Alk | ||
| Mouse | Plasmid DNA | Pten, p53, β-catenin | ||
| Mouse | AAV | NeUN | ||
| Mouse | Adenovirus | Cebpa | ||
| 3 | Lymphoma | Mouse | Lentivirus | Tet2, NF1, EZH2,RUNX1 |
| Mouse | Lentivirus | Mcl, p53 | ||
| Mouse | Retrovirus | p53 | ||
| Mouse | DNA electroporaion | Mll3 |
Tumor Modeling Via CRISPR-Cas System:
Converting a normal cell into a tumor cell is a multistep procedure in which the cell must go through a number of mutations [19, 20]. Scientists were successful in modeling tumor cells by inducing mutation at great speed and precision using the CRISPR/Cas9 system. Some studies reported that they have successfully applied this technique to model liver cancer cells by inducing mutations in cancer associated genes in vivo.
Successfully truncating the APC tumor suppressor gene in gut cells was indicative of a well-developed early event in the genesis of colorectal cancer. In order to choose APC-deficient human intestinal stem cells, the Wnt signaling activators have been removed from the growth media. This has induced β-catenin stability and up-regulation of the Wnt pathway [21, 22]. Other methods involve producing cells that have both a loss-of-function mutation and the KRAS oncogene active. Additionally, CRISPR/Cas9 was used to deactivate P53 [23]. These methods might also be used to create a mixture of mutations, which could then be investigated by injecting these modified cells into immunodeficient mice. The big question of whether the cancer-causing mutations happen randomly or in a specific order of time is answered by this cancer model.
Similar techniques have been utilized to use CRISPR/ Cas9, which targets multiple protein-coding genes as well as miRNA precursors, to transform a non-metastatic mouse lung tumor cell line. After injecting the transformed cells into immunodeficient mice, it was observed a large increase in tumor and lung metastasis [24]. The scientists used deep sequencing to identify numerous new genes whose expression was essential for tumor development and metastasis. This method enables the recapitulation of the development and metastasis of tumors, which may aid in the development of particular treatments that specifically target defective genes [25].
CRISPR/Cas9-mediated targeting human papillomavirus E6
The RNA guided endonuclease CRISPR-Cas9 is developed from the bacteria Streptococcus pyogenes. It has been widely employed for genome engineering in a range of species, as well as the genus plasmodium Leishmania, the underlying cause of human leishmaniasis, because of its simplicity, adaptability, and high effectiveness. CRISPR- Cas9 has been demonstrated to be a more efficient tool for deleting or disrupting Leishmania genes, generating point mutations, and adding tags to endogenous genes than classical homologous recombination gene targeting. The consistent CRISPR expression systems were demonstrated to eliminate multigene family Leishmania genes as well as genes contained in multiploid chromosomes, identify the critical Leishmania genes, and construct particular chromosomal translocations [26]. Cervical cancer caused by the human papillomavirus (HPV) is a serious health concern for women in developing countries. As its diagnosis and poor prognosis, it has a significant fatality rate. The development and major factor that leads to this kind of cancer are entirely dependent on two main oncogenes, E6 and E7, which are constitutively expressed, leading to carcinogenesis [27].
Human papillomaviruses of the Papillomaviridae family are a kind of tiny non-enveloped circular double-stranded DNA virus that measures 50-55 nm in diameter. The category has 300 distinct genotypes, 200 of which are believed to be harmful to humans. The scientists categorized papillomaviruses into genera, species, kinds, and subtypes based on gene sequences similarity of the L1 ORFs of 118 papillomavirus types (Pal & Kundu, 2020). Human papillomaviruses have been classified into five genera:
1. alpha (65 types including HPV16, 18, 31, 33, etc.)
2. beta (53 types including HPV5, 9, 49, etc.
3. gamma (98 types including HPV4, 48, 50, etc.)
4. mu (3 types including HPV1, HPV63, and HPV 204)
5. nu (HPV41). Among them, alpha-papillomaviruses are the most commonly focused group of papillomaviruses since they are known to be responsible for 5% of cancer occurrences worldwide [27].
The fourth most frequent cancer among women globally is cervical cancer [28]. The most prevalent high- risk HPV type and the one that carries the highest risk of developing cervical cancer is human papillomavirus type 16 (HPV16). In their lifetime, over than 80 percent of women who had at least one partner of the other sex will get HPV [29]. Persistent high-risk HPV infection has been identified as a major factor in the development of cervical cancer. Additionally, it was discovered by researchers that the HPV viruses can incorporate their DNA into the human genome, which appeared to be a crucial step in the development of carcinogenesis [30]. The HPV oncogene is persistently expressed as a result of integration, making it challenging to eradicate. For individuals with a chronic HPV infection and the incorporation of HPV genes, there is currently no viable therapy. Zinc finger nuclease (ZFN), transcription activator-like effectors nuclease (TALEN), and clustered regularly interspaced short palindrome repeat (CRISPR/Cas9) are the three primary gene-editing techniques. All of these techniques for gene editing may cause DNA double-strand breaks (DSBs) that were specifically targeted and modify genes by triggering DNA repair processes. Gene therapy is getting more accurate and efficient with the advancement of these gene-editing techniques [31]. These gene-editing strategies made targeting HPV oncoprotein genes might have a significant impact on the targeted cells. The effectiveness of gene treatments in treating HPV infection sickness has not yet been compared. Researchers just need to create the gRNA complementary to such target DNA sequence for the CRISPR/Cas9 system; no further components are required. Because it is quick and simple to create, the CRISPR/Cas9 system may be an appropriate substitute for ZFN and TALEN for initiating targeted gene editing. It might result in gene deletion, reversion, and insertion by causing DNA damage at a specific spot, which could then be repaired by the cell’s self-repairing machinery by NHEJ (non-homologous end-joining) or HDR (homologous-dependent repair). According to some earlier research, the HPV oncogene targeted by the CRISPR/Cas9 system may be able to treat HPV-induced cervical cancer. However, there hasn’t been much dynamic monitoring of the therapy process up to this point in the suitable animal model.
Protein degradation in breast cancer
A type of cancer that originates in breast cells is called breast cancer. Some of the signs and symptoms of breast cancer include:
• A newly reversed nipple
• A breast lump or thickening that feels different from the surrounding tissue
• A change in a breast’s size, shape, or appearance
• Variations in the skin covering the breast, such as dimpling;
• Peeling, scaling, crusting, or flaking of the pigmented place of skin around the nipple (areola) or breast skin;
• A newly inverted nipple; [32]
Both the rates of synthesis and the rates of degradation affect the number of proteins within cells. Asymmetric rates of protein breakdown are a key component of cell control. The ½ of proteins inside cells can range greatly, from a few minutes to many days. Many proteins that degrade quickly serve as regulatory molecules, including transcription factors. These proteins must have a quick turnover in order for their levels to react swiftly to outside stimuli. Another way for controlling the activity of intracellular enzymes is provided by the fast degradation of other proteins in response to certain signals. Furthermore, defective or damaged proteins are identified and quickly destroyed within cells, eradicating the effects of errors produced during protein synthesis. Protein breakdown in eukaryotic cells is mediated by two main pathways: lysosomal proteolysis and the ubiquitin-proteasome system.
The enhanced action of protein degradation is one of the key factors in promoting tumor cell growth. The 26S proteasome, an enzyme that is important for protein degradation including cell cycle control and apoptosis-related proteins, is one of these protein-degrading enzymes [33, 34]. Proteasome inhibitors have been shown to have anticancer and apoptosis-enhancing effects in cancer model organisms. Additionally, it makes tumor cells more susceptible to both intrinsic and extrinsic pro-apoptotic signals. As a result, the proteasome is now a target for cancer therapies. There is evidence that a site-specific protease phosphorylation mechanism regulates the growth of breast cancer [35], suggesting that disrupting and interfering with this mechanism might be useful in containing the illness. To stop the carcinogenesis of mice with proteasome-dependent human breast cancer cells, dual-specificity tyrosine-regulated kinase 2 (DYRK2) deletion (the proteins that phosphorylate the proteasome components) was developed [36]. Treatment options for ER-positive breast cancer include utilizing aromatase inhibitors (AI) to prevent the production of estrogen, as well as tamoxifen and fulvestrant, which compete with estrogen for ER. AI and tamoxifen are ineffective against advanced metastatic breast carcinoma caused by ER mutations like ERY537S and ERD538G [37]. ERY537S or ERD538G was substituted for the wild-type form of ER in a breast cancer positive ER model using CRISPR/Cas9 to demonstrate the impact of these mutations. These mutant cells exhibit estrogen independent behavior and are somewhat resistant to antiestrogen. It was discovered that the mutant cells’ antiestrogen resistance was connected to an increase in the polypeptide response that lessens ER degradation [38]. Breast cancer metastasis and cancer progression are influenced by migration and invasion enhancers (MIEN1). According to research, increasing MIEN1 expression may promote tumor spread and migration. Through the use of CRISPR/Cas9, a specific removal in this gene successfully reversed its expression, therefore controlling the disease’s progression. This method enables us to comprehend the part MIEN1 plays in the development of cancer and tumors in great detail, which may one day lead to the development of a breast cancer treatment option [39]. PTEN (phosphatase and tensin homolog) gene mutations are a critical stage in the development of cancer. PTEN is a tumor suppressor gene that regulates the cell cycle and regulates the rate of proliferation. In order to disrupt PTEN in mice with mammary gland-specific lack of E-cadherin, invasive lobular breast cancer (ILC)-initiating cells were specifically targeted using CRISPR/Cas9. This method may be used to quickly test in vivo potential tumor suppressor genes. ILC [40].
Use of CRISPR cas9 system for countermanding drugs resistance in cervical cancer:
Chemotherapy is currently the most viable cervical cancer treatment. A significant obstacle in cervical cancer treatment is cellular resistance, which develops as a result of the prolonged use of chemotherapeutic medications. The human papillomavirus (HPV) transmission, which results in the integration of the virus’ oncogenes E6 and E7 into host cells, is the primary cause of the significant number of cervical malignancies [41]. Therefore, the element for inducing and retaining cervical cancerization is the over expression of these two genes [42]. They provide the novelty and security of the nanoscale editing method as the HPV-derived E6 and E7 and are present in the cells of cervical cancer only [43].
The chemotherapeutic drug Docetaxel (DOC) and the CRISPR/Cas9 system were employed in conjunction to overcome the resistance to transformational therapy and enhance the therapeutic effect. In the cervical cancer model of the mouse, the nano-system created by encapsulating hydrophobic DOC negatively charged CRISPR/Cas9 plasmid, and cationic liposomes demonstrated great therapeutic efficacy and few adverse effects [44]. In order to prevent cervical cancer from growing and spreading from the start, the E6 and E7 oncogenes deletion is intended. The attacking of the E7 gene of HPV increased the expression level of Rb protein, which was found to increase 2.58-fold and 2.42-fold in expression. HPV E6 protein processes suppress the p53 tumor suppressor expression of the host, so the decline of E6 activity is anticipated to end in the stimulation of the expression of p53. Chemo resistance is significantly influenced by pRB and p53. After eliminating the HeLa cell’s E6 and E7 genes, the CRISPR system increased the pRB and p53 gene expression, which turns to induce HeLa cells to undergo apoptosis [45]. Some CRISPR approaches along with their target genes are being represented in Table 2.
| Target Genes | Cell line | CRISPR approach | References |
| FASN | MCF-7 | Type 2 CRISPR/Cas9 | (Gonzalez-Salinas et al., 2020) [54] |
| PTEN | SUM159 | CRISPR activation | (Choudhury et al., 2016) [55] |
| CDK7 | TNBC | CRISPR/Cas9 genetic editing | (Y. Wang et al., 2015) [56] |
| MASTL | Human Mammary tumor cell lines | CRISPR based interruption | (Álvarez-Fernández et al., 2018) [57] |
| MYC oncogene | _ | CRISPR/Cas9 mediated mutagenesis | (Schuijers et al., 2018) [58] |
| CXCR7, CXCR4 | MDA-MB231 | CRISPR/Cas9 knockout | (S. Luo et al., 2016) [59] |
The DNA endonucleases guided by RNA were developed by introducing the sgRNAs that were HPV specific into the px330 vector that was expressing S. pyogenes Cas9 to produce the px330-E6E7 plasmid, also known as E6E7, and were proven to be a successful approach. In HPV-positive cervical carcinoma cells, disruption of E6E7 led to apoptosis induction and tumor growth suppression [46]. Furthermore, it was revealed that combined treatment reduced the dosage of docetaxel used, and the CRISPR system’s attack decreased the adverse impact on other organs. DOC impact absorption into cells occurs significantly more quickly than CRISPR/ Cas9. Therefore, it would be advantageous to combine CRISPR therapy with conventional chemotherapy, so that the challenge of drug resistance and tumor inhibition be efficiently treated (Figure 3).
Figure 3. CRISPR cas9 in Reversing Drug Resistance in Cervical Cancer.
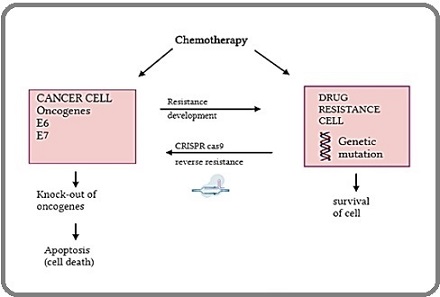
Kinesin-5 A133P mutation in ispinesib resistance in cervical cancer
A molecular motor protein called kinesin-5 is crucial for mitosis. An emerging body of research has revealed that kinesin-5 is involved in the development of tumors and is seen as a possible target for cervical cancer therapy [15]. Some of the molecules involved in resistance to cancer and their effects are briefly shown in Table 3.
| Therapeutic agents | Molecules involved in resistance | Cancer | Results |
| Cytotoxic agents | |||
| Doxorubicin | P-glycoprotein | Breast cancer | Increases sensitivity to doxorubicin. |
| Epirubicin | MLL | Bladder cancer | Reverse drug resistance. |
| Cisplatin | p53, CTR | Oesophageal adenocarcinoma | Inhibits cell growth and induces cell cycle arrest. |
| Paclitaxel | Rsf-1 | Lung cancer | Reverse drug resistance. |
| Immunotherapy | |||
| Cancer vaccines and adaptive T cell therapies | PBAF | Melanoma | Increases sensitivity to immunotherapy. |
| Molecular targeted agents | |||
| Ispinesib | Kinesin-5 A133P | Cervical cancer | Overthrows the previous resistance mechanism. |
| Imatinib | ASXL1 | Chronicmyelocytic leukemia | Enhances differentiation ability. |
| Bortezomib | Rpn13 | Multiple myeloma | Inhibits proliferation. |
| Trastuzumab | HER2 | Breast cancer | Reverse drug resistance. |
| SAHA | p57 | Pancreatic ductal adenocarcinoma | Reverse drug resistance. |
Ispinesib, a kinesin-5 inhibitor, has started to be tested in human clinical trials as a cancer treatment. The over expression of kinesin-12 or activation of the EGFR, which were previously thought to be resistance mechanisms, did not contribute to Ispinesib resistance, according to a recent CRISPR-Cas9-based screening investigation.
CRISPR cas9 applications in cervical carcinogenesis
The abnormal proliferation of cells on the lining of cervix is what causes cervical cancer. In the type of cancers in women worldwide this type is the fourth most frequently occurring cancer. The most pervasive high-risk HPV type and the one that carries the highest risk of developing cervical cancer is human papillomavirus type 16 (HPV16). In their lifetime, more than 80% of women who have had at least one partner of the other sex will contract HPV [47]. It has very long been thought to be a major factor in the development of cervical cancer is because of high-risk HPV infection. The screening methods for cervical cancer shown by pap smears and the testing of high-risk HPV has shown some advances in recent years [48]. In most cases, surgical interventions were considered the prominent treatment for cervical cancer patients, but these had side effects on the patients and could also have a promising effect on the pregnancy of women for an extended period of time.
CRISPR cas9 system has proved to have favorable effects on cervical pre-cancer treatment and to the already present treatments of infections of cervix related to HPV, considered a promising restorative strategy [49]. The human papillomavirus (HPV) infection, which results in the integration of the virus’ oncogenes E6 and E7 into host cells, is the leading cause of cervical cancer. Therefore, the key to causing and maintaining cervical cell cancerization is the over expression of these two genes [50]. Many types of research have confirmed the role of HPV E6 and E7 oncogenes targeted by CRISPR cas system. While the E7 protein was genetically conserved, the HPV16 genome shared several amino acid-changing variations. Additionally, the E7 HPV16 is thought as the only oncoprotein that has the potential to immortalize human keratinocytes in vitro and induce cervical cancer in an animal model. It follows that HPV16 E7 that is targeted by CRISPR cas system may had interesting therapeutic applications (Figure 4).
Figure 4. Applications of CRISPR cas9 in Treatment of Cervical Carcinogenesis.
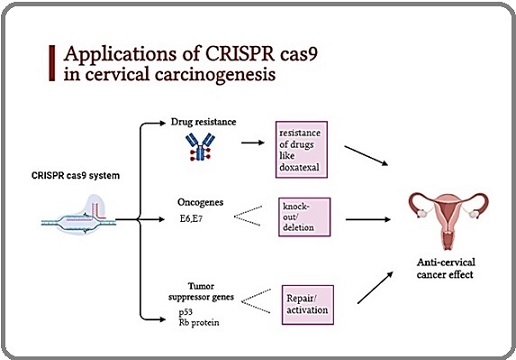
By this HPV16 E7 is considered as a suitable target for the HPV16-induced cervical cancer therapy. Additionally, in vitro and in naked mice models, gene therapy has shown encouraging effects, and theE6 gene of HPV16 is a strong aspirant cleavage site for this treatment.
Some of the gene editing tools like ZFN and TALENS have also shown the same efficacy as the CRISPR system in the treatment of cervical cancer [51]. The TALEN and ZFN are secluded proteins that have the flexible DNA restricting domain intertwined with FokI, that is an intricate design procedure for these tools to target a unique DNA sequence. The CRISPR system, in contrast, is reasonably straightforward method created to address this deficiency [52]. Additionally, because shRNA primarily target the products of transcription of the HPV16 oncogene, it had little effect on the HPV DNA that had been incorporated in the cells of human, indicating that such strategy could not be long lasting. Additionally, the CRISPR/Cas9 system may edit many genes simultaneously by creating different sgRNAs that attack various genomic sequences, which may significantly increase the technology’s therapeutic potential [53]. Therefore, it is believed that however, in some cell lines CRISPR along with some other gene editing tools has the same efficacy; it still is the more promising tool for the therapy of cervical lesions.
Future perspectives
The CRISPR cas9 system is thought to be the most useful for clinical protocols. Since it is discovered, CRISPR system has made it possible to investigate the function of specific genes in intricate biological processes. Technologies for CRISPR/Cas9-based screening have advanced, raising hopes for novel findings. Clinical trials for CRISPR/Cas9-based treatments, however, have been hampered by several issues, such as the possibility of off-target effects, the need to optimize attack sites, safety concerns while working in vivo, and a lack of available drug delivery options. For instance, techniques must be improved to transport the components of CRISPR/Cas9 safely and effectively in the targeted tissues. This genome editing technique has the potential to be dependable as well as simple to use. CRISPR/Cas9 has shown many ways for unveiling gene function in cancer treatment thanks to its simplicity and versatility. For example, current findings from genome-wide deep sequencing will be useful for choosing appropriate target sites and creating greatly specified gRNA. Additionally, interactions between CRISPR/Cas9 and additional chemo/radio therapeutic drugs may open new therapy options and enhance the clinical outcomes of cervical and breast cancer patients.
In conclusion, since scientists discovered they could use CRISPR to alter the DNA of any animal easily and precisely, it has revolutionized biology. It is a potent instrument that could be utilized to change the in a way that future generations could inherit the genome. Since its discovery, CRISPR/Cas9 has made it possible to investigate the function of specific genes in intricate biological processes. Technologies for CRISPR/Cas9-based screening have advanced, raising hopes for novel findings. For application in therapeutic studies; the CRISPR/Cas9 system enables precise editing of a target sequence in model organisms and humans. Theoretically, it is also conceivable to treat malignancies as well as genetic and viral illnesses. For cancer patients, the expansion of full genome libraries is made possible by CRISPR/Cas9, a very adaptable and practical method. CRISPR cas9 has proved to be the novel therapeutic strategy for the treatment of breast cancer and cervical cancer. Clinical results for patients with cervical and breast cancer may be improved by the interactions and alliances between CRISPR/Cas9 and other chemo/radio therapeutic medicines.
Conflict of interest
None
Funding
None
Ethical approval
Not applicable
References
- CRISPR-Cas9: a promising genetic engineering approach in cancer research Ratan ZA , Son Y, Haidere MF , Uddin BMM , Yusuf MA , Zaman SB , Kim J, Banu LA , Cho JY . Therapeutic Advances in Medical Oncology.2018;10. CrossRef
- Advances in CRISPR-Cas based genome engineering Katrekar D, Hu M, Mali P. Current Opinion in Biomedical Engineering.2017;1. CrossRef
- Immunity to CRISPR Cas9 and Cas12a therapeutics Chew WL . Wiley Interdisciplinary Reviews. Systems Biology and Medicine.2018;10(1). CrossRef
- Involvement of Human Papillomaviruses in Cervical Cancer Wang X, Huang Xi, Zhang Y. Frontiers in Microbiology.2018;9. CrossRef
- Breast cancer statistics, 2017, racial disparity in mortality by state DeSantis CE , Ma J, Goding Sauer A, Newman LA , Jemal A. CA: a cancer journal for clinicians.2017;67(6). CrossRef
- DNA Methylation Patterns in Normal Tissue Correlate more Strongly with Breast Cancer Status than Copy-Number Variants Gao Y, Widschwendter M, Teschendorff AE . EBioMedicine.2018;31. CrossRef
- New drugs for breast cancer subtypes: targeting driver pathways to overcome resistance Curigliano G. Cancer Treatment Reviews.2012;38(4). CrossRef
- Upregulation of ER Signaling as an Adaptive Mechanism of Cell Survival in HER2-Positive Breast Tumors Treated with Anti-HER2 Therapy Giuliano M, Hu H, Wang Y, Fu X, Nardone A, Herrera S, Mao S, et al . Clinical Cancer Research: An Official Journal of the American Association for Cancer Research.2015;21(17). CrossRef
- Understanding and treating triple-negative breast cancer Anders C, Carey LA . Oncology (Williston Park, N.Y.).2008;22(11).
- Two hundred years of cancer research DeVita VT , Rosenberg SA . The New England Journal of Medicine.2012;366(23). CrossRef
- Targeting tyrosine-kinases and estrogen receptor abrogates resistance to endocrine therapy in breast cancer Liu S, Meng X, Chen H, Liu W, Miller T, Murph M, Lu Yi, et al . Oncotarget.2014;5(19). CrossRef
- Applications of the CRISPR/Cas9 system in murine cancer modeling Zuckermann M, Kawauchi D, Gronych J. Briefings in Functional Genomics.2017;16(1). CrossRef
- A CRISPR view of gene regulation Banerjee B, Sherwood RI . Current Opinion in Systems Biology.2017;1. CrossRef
- Programmable sequential mutagenesis by inducible Cpf1 crRNA array inversion Chow RD , Kim HR , Chen S. Nature Communications.2018;9(1). CrossRef
- The CRISPR-Cas9 system: a promising tool for discovering potential approaches to overcome drug resistance in cancer Zhang J, Zhou W, Wang X, Wang L. RSC advances.2018;8(58). CrossRef
- Use of CRISPR/Cas9 gene-editing tools for developing models in drug discovery Ahmad G, Amiji M. Drug Discovery Today.2018;23(3). CrossRef
- HIT-Cas9: A CRISPR/Cas9 Genome-Editing Device under Tight and Effective Drug Control Zhao C, Zhao Y, Zhang J, Lu J, Chen L, Zhang Y, Ying Y, et al . Molecular Therapy. Nucleic Acids.2018;13. CrossRef
- Bacterial genetics: What CRISPR memories are made of Du Toit A. Nature Reviews. Microbiology.2015;13(4). CrossRef
- Age at cancer diagnosis and interpretation of survival statistics Autier P. The Lancet. Oncology.2016;17(7). CrossRef
- Break Breast Cancer Addiction by CRISPR/Cas9 Genome Editing Yang H, Jaeger M, Walker A, Wei D, Leiker K, Weitao T. Journal of Cancer.2018;9(2). CrossRef
- Use of CRISPR-modified human stem cell organoids to study the origin of mutational signatures in cancer Drost J, Boxtel R, Blokzijl F, Mizutani T, Sasaki N, Sasselli V, Ligt J, et al . Science (New York, N.Y.).2017;358(6360). CrossRef
- Modeling colorectal cancer using CRISPR-Cas9-mediated engineering of human intestinal organoids Matano M, Date S, Shimokawa M, Takano A, Fujii M, Ohta Y, Watanabe T, Kanai T, Sato T. Nature Medicine.2015;21(3). CrossRef
- Targeting mutant KRAS with CRISPR-Cas9 controls tumor growth Wonjoo K, Sangeun L, Han Sang K, Minjung S, Yong Hoon C, Young-Hoon K, Jeonghong S, et al . Genome Research.2018;28(3). CrossRef
- Combined use of EpCAM and FRα enables the high-efficiency capture of circulating tumor cells in non-small cell lung cancer Chen L, Peng M, Li N, Song Q, Yao Y, Xu B, Liu H, Ruan P. Scientific Reports.2018;8(1). CrossRef
- Genome-wide CRISPR screen in a mouse model of tumor growth and metastasis Chen S, Sanjana NE , Zheng K, Shalem O, Lee K, Shi X, Scott DA , et al . Cell.2015;160(6). CrossRef
- Genetic editing and interrogation with Cpf1 and caged truncated pre-tRNA-like crRNA in mammalian cells Zhang X, Xu L, Fan R, Gao Q, Song Y, Lyu X, Ren J, Song Y. Cell Discovery.2018;4. CrossRef
- Human Papillomavirus E6 and E7: The Cervical Cancer Hallmarks and Targets for Therapy Pal A, Kundu R. Frontiers in Microbiology.2019;10. CrossRef
- Prevalence of Cervical Cancer and Associated Factors Among Women Attended Cervical Cancer Screening Center at Gahandi Memorial Hospital, Ethiopia Mekuria M, Edosa K, Endashaw M, Bala ET , Chaka EE , Deriba BS , Tesfa B. Cancer Informatics.2021;20. CrossRef
- Human papillomaviruses Korsman SNJ , van Zyl GU , Nutt L , Andersson MI , Preiser W . Virology, 66–67. https://doi.org/(تو نت پیدا نشد).2012;:66-67. CrossRef
- Nonviral gene editing via CRISPR/Cas9 delivery by membrane-disruptive and endosomolytic helical polypeptide Wang H, Song Z, Lao Y, Xu X, Gong J, Cheng D, Chakraborty S, et al . Proceedings of the National Academy of Sciences of the United States of America.2018;115(19). CrossRef
- Applications of genome editing technology in the targeted therapy of human diseases: mechanisms, advances and prospects Li H, Yang Y, Hong W, Huang M, Wu M, Zhao X. Signal Transduction and Targeted Therapy.2020;5(1). CrossRef
- Breast cancer. https://my.clevelandclinic.org/health/diseases/3986-breast-cancer Cleveland Clinic. . 2022.
- Ancient drug curcumin impedes 26S proteasome activity by direct inhibition of dual-specificity tyrosine-regulated kinase 2 Banerjee S, Ji C, Mayfield JE , Goel A, Xiao J, Dixon JE , Guo X. Proceedings of the National Academy of Sciences of the United States of America.2018;115(32). CrossRef
- TRIM11 activates the proteasome and promotes overall protein degradation by regulating USP14 Chen L, Zhu G, Johns EM , Yang X. Nature Communications.2018;9(1). CrossRef
- A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA , Charpentier E. Science (New York, N.Y.).2012;337(6096). CrossRef
- Site-specific proteasome phosphorylation controls cell proliferation and tumorigenesis Guo X, Wang X, Wang Z, Banerjee S, Yang J, Huang L, Dixon JE . Nature Cell Biology.2016;18(2). CrossRef
- A Computational Assay of Estrogen Receptor α Antagonists Reveals the Key Common Structural Traits of Drugs Effectively Fighting Refractory Breast Cancers Pavlin M, Spinello A, Pennati M, Zaffaroni N, Gobbi S, Bisi A, Colombo G, Magistrato A. Scientific Reports.2018;8(1). CrossRef
- Role of Brf1 interaction with ERα, and significance of its overexpression, in human breast cancer Fang Z, Yi Y, Shi G, Li S, Chen S, Lin Y, Li Z, He Z, Li W, Zhong S. Molecular Oncology.2017;11(12). CrossRef
- CRISPR deletion of MIEN1 in breast cancer cells Van Treuren T, Vishwanatha JK . PloS One.2018;13(10). CrossRef
- Modeling invasive lobular breast carcinoma by CRISPR/Cas9-mediated somatic genome editing of the mammary gland Annunziato S, Kas SM , Nethe M, Yücel H, Del Bravo J, Pritchard C, Bin Ali R, et al . Genes & Development.2016;30(12). CrossRef
- The natural history of cervical HPV infection: unresolved issues Woodman CBJ , Collins SI , Young LS . Nature Reviews. Cancer.2007;7(1). CrossRef
- Inactivation of the human papillomavirus E6 or E7 gene in cervical carcinoma cells by using a bacterial CRISPR/Cas RNA-guided endonuclease Kennedy EM , Kornepati AVR , Goldstein M, Bogerd HP , Poling BC , Whisnant AW , Kastan MB , Cullen BR . Journal of Virology.2014;88(20). CrossRef
- Prevalence and genotype distribution of HPV and cervical pathological results in Sichuan Province, China: a three years surveys prior to mass HPV vaccination Luo Q, Jiang N, Wu Q, Wang J, Zhong J. Virology Journal.2020;17(1). CrossRef
- The vaginal microbiota, human papillomavirus and cervical dysplasia: a systematic review and network meta-analysis Norenhag J, Du J, Olovsson M, Verstraelen H, Engstrand L, Brusselaers N. BJOG: an international journal of obstetrics and gynaecology.2020;127(2). CrossRef
- Epidemiologic classification of human papillomavirus types associated with cervical cancer Muñoz N, Bosch FX , Sanjosé S, Herrero R, Castellsagué X, Shah KV , Snijders PJF , Meijer CJLM . The New England Journal of Medicine.2003;348(6). CrossRef
- Disruption of HPV16-E7 by CRISPR/Cas system induces apoptosis and growth inhibition in HPV16 positive human cervical cancer cells Hu Z, Yu L, Zhu D, Ding W, Wang X, Zhang C, Wang L, et al . BioMed Research International.2014;2014. CrossRef
- HPV16 E7 Genetic Conservation Is Critical to Carcinogenesis Mirabello L, Yeager M, Yu K, Clifford GM , Xiao Y, Zhu B, Cullen M, et al . Cell.2017;170(6). CrossRef
- The application of CRISPR/Cas9 system in cervical carcinogenesis Gao C, Wu P, Yu L, Liu L, Liu H, Tan X, Wang L, Huang X, Wang H. Cancer Gene Therapy.2022;29(5). CrossRef
- The E7 gene of human papillomavirus type 16 is sufficient for immortalization of human epithelial cells Halbert CL , Demers GW , Galloway DA . Journal of Virology.1991;65(1). CrossRef
- Oncogenic Human Papillomavirus: Application of CRISPR/Cas9 Therapeutic Strategies for Cervical Cancer Zhen S, Li X. Cellular Physiology and Biochemistry: International Journal of Experimental Cellular Physiology, Biochemistry, and Pharmacology.2017;44(6). CrossRef
- Application of new biotechnologies for improvements in swine nutrition and pork production Wu G, Bazer FW . Journal of Animal Science and Biotechnology.2019;10. CrossRef
- CRISPR/Cas9 for genome editing: progress, implications and challenges Zhang F, Wen Y, Guo X. Human Molecular Genetics.2014;23(R1). CrossRef
- CRISPR-Cas9: a new and promising player in gene therapy Xiao-Jie L, Hui-Ying X, Zun-Ping K, Jin-Lian C, Li-Juan J. Journal of Medical Genetics.2015;52(5). CrossRef
- Transcriptomic and cellular analyses of CRISPR/Cas9-mediated edition of FASN show inhibition of aggressive characteristics in breast cancer cells Gonzalez-Salinas F, Rojo R, Martinez-Amador C, Herrera-Gamboa J, Trevino V. Biochemical and Biophysical Research Communications.2020;529(2). CrossRef
- CRISPR-dCas9 mediated TET1 targeting for selective DNA demethylation at BRCA1 promoter Choudhury SR , Cui Y, Lubecka K, Stefanska B, Irudayaraj J. Oncotarget.2016;7(29). CrossRef
- CDK7-dependent transcriptional addiction in triple-negative breast cancer Wang Y, Zhang T, Kwiatkowski N, Abraham BJ , Lee TJ , Xie S, Yuzugullu H, et al . Cell.2015;163(1). CrossRef
- Therapeutic relevance of the PP2A-B55 inhibitory kinase MASTL/Greatwall in breast cancer Álvarez-Fernández M, Sanz-Flores M, Sanz-Castillo B, Salazar-Roa M, Partida D, Zapatero-Solana E, Ali HR , et al . Cell Death and Differentiation.2018;25(5). CrossRef
- Transcriptional Dysregulation of MYC Reveals Common Enhancer-Docking Mechanism Schuijers J, Manteiga JC , Weintraub AS , Day DS , Zamudio AV , Hnisz D, Lee TI , Young RA . Cell Reports.2018;23(2). CrossRef
- The association of PTEN hypermethylation and breast cancer: a meta-analysis Luo S, Chen J, Mo X. OncoTargets and Therapy.2016;9. CrossRef
Author Details
How to Cite
- Abstract viewed - 0 times
- PDF (FULL TEXT) downloaded - 0 times
- XML downloaded - 0 times